We offer fully funded PhD and postdoc positions.
Statistical physics of living systems
Welcome to the Statistical Physics of Living Systems group
Biological systems rely on an influx of energy from the environment to build and maintain complex patio-temporal structures. Striking examples of the non-equilibrium nature of biological systems are the self-organisation of cells during embryonic development, regeneration and tumour genesis. With recent technological breakthroughs in imaging and genomics we are now able to probe these processes with unprecedented microscopic detail. Biological function, however, relies on collective degrees of freedom that are not straight forwardly related to microscopic states. In our group we combine novel technologies developed and used by our experimental collaborators with tools from non-equilibrium statistical physics to understand collective processes underlying the behaviour of active biosystems. Understanding the mechanistic principles underlying these systems is not only pivotal for the development of regenerative therapies, but also gives rise to challenging questions at the frontier of non-equilibrium physics. Read more about our research.
***Please note that as of Oct 1 the group has moved to the Ludwig-Maximilians Universität München in Munich.***
The Statistical Physics of Living Systems group was started in 2017 and is embedded into the Biological Physics division of the Max Planck Institute for the Physics of Complex Systems and the Center for Systems Biology in Dresden (Germany). The group maintains close collaborations with experimental groups on the local and international level.
Funding



Research highlights
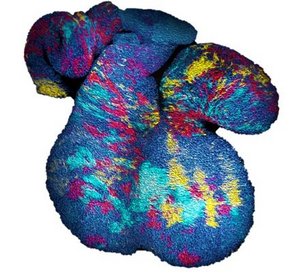
Scaling and universality of clone dynamics during tissue development
Despite the complexity of organ development, clonal dynamics may converge to a critical state characterized by universal scaling behaviour of clone sizes. Our work allows to identify experimental strategies that unveil cell fate behaviour during development and tumour growth.
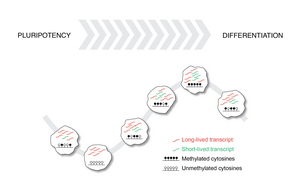
Epigenetic dynamics during embryonic development
DNA methylation is the primary layer of modifications of the DNA. We found that during the early stages of embryonic development these epigenetic marks undergo collective (genome-wide) oscillations. Our work shows that epigenetic changes can be surprisingly dynamic and it highlights how mechanistic understanding of cellular decision making can be obtained by combining single-cell genomics with non-equilibrium physics.
Design principles of pattern forming systems
What are the molecular mechanisms of robust embryonic patterning? We studied design principle of gene regulatory networks subject to intrinsic noise and extrinsic perturbations.